Functionally motivated dimensionality reduction methods for neural population dynamics
 Brain functions, ranging from perception to cognition and action, are produced by the collective dynamics of populations of neurons. High-dimensional neural population data are commonly analyzed in lower-dimensional subspaces, which enables characterization of dynamic neural phenomena. However, subspaces themselves do not provide a principled explanation of those phenomena. In particular, the vast majority of dimensionality reduction methods in computational neuroscience attempt to account for variance in the data through latent factors. This approach has two principal shortcomings. First, by treating the observed neural activity as simply a readout of latent dynamics, the role of the neurons (i.e. the physical degrees of freedom of the system) in producing the observed dynamics is obscured. Second, the objective of maximizing variance within the reduced dimensional space presupposes that it is high variance activity that carries out computations of interest (which may or may not be true). To address these shortcomings, we have developed several dimensionality reduction methods that identify latent subspaces via interpretable embeddings (e.g., linear projections) of the observed neural activity by optimizing objective functions that directly encode functional measures of interest. Examples of the latter include dynamical complexity (Dynamical Components Analysis) and feedback controllability (Feedback Controllable Components Analysis). Furthermore, we are actively developing methods that consider simultaneous, functionally relevant dimensionality reduction across multiple co-recorded time series (e.g. multiple brain areas or high dimensional behavioral or stimulus time series). These methods enable drawing direct connections between low-dimensional dynamics, behaviorally relevant computations, and the underlying populations of neurons.
Brain functions, ranging from perception to cognition and action, are produced by the collective dynamics of populations of neurons. High-dimensional neural population data are commonly analyzed in lower-dimensional subspaces, which enables characterization of dynamic neural phenomena. However, subspaces themselves do not provide a principled explanation of those phenomena. In particular, the vast majority of dimensionality reduction methods in computational neuroscience attempt to account for variance in the data through latent factors. This approach has two principal shortcomings. First, by treating the observed neural activity as simply a readout of latent dynamics, the role of the neurons (i.e. the physical degrees of freedom of the system) in producing the observed dynamics is obscured. Second, the objective of maximizing variance within the reduced dimensional space presupposes that it is high variance activity that carries out computations of interest (which may or may not be true). To address these shortcomings, we have developed several dimensionality reduction methods that identify latent subspaces via interpretable embeddings (e.g., linear projections) of the observed neural activity by optimizing objective functions that directly encode functional measures of interest. Examples of the latter include dynamical complexity (Dynamical Components Analysis) and feedback controllability (Feedback Controllable Components Analysis). Furthermore, we are actively developing methods that consider simultaneous, functionally relevant dimensionality reduction across multiple co-recorded time series (e.g. multiple brain areas or high dimensional behavioral or stimulus time series). These methods enable drawing direct connections between low-dimensional dynamics, behaviorally relevant computations, and the underlying populations of neurons.
Data-driven identification of functionally relevant wiring principles from connectomics
 Modern large scale connectomics is producing increasingly detailed datasets of the ground truth connectivity between neurons across model species. A key open challenge is the development of theory and statistical techniques that can leverage these data to produce novel insights into the relationship between structure and function. Prior work on identifying the wiring principles that constitute neuronal networks have uncovered several robust observations, including structured connectivity between genetically defined cell-types, tradeoffs between total wiring cost and communication efficiency, and an overabundance of higher order network motifs relative to unstructured, random networks. Statistically, these observations can be encoded into models of random graphs inspired by statistical mechanics that match the connectivity patterns of observed networks at various spatial scales. These models can then be used as generative distributions to interrogate the specific structural mechanisms and design tradeoffs that enable particular functions (e.g. control of network dynamics). We are actively developing methods to fit and sample from these generative models at the scales needed to bring insight into the controllability properties of real-world connectomes.
Modern large scale connectomics is producing increasingly detailed datasets of the ground truth connectivity between neurons across model species. A key open challenge is the development of theory and statistical techniques that can leverage these data to produce novel insights into the relationship between structure and function. Prior work on identifying the wiring principles that constitute neuronal networks have uncovered several robust observations, including structured connectivity between genetically defined cell-types, tradeoffs between total wiring cost and communication efficiency, and an overabundance of higher order network motifs relative to unstructured, random networks. Statistically, these observations can be encoded into models of random graphs inspired by statistical mechanics that match the connectivity patterns of observed networks at various spatial scales. These models can then be used as generative distributions to interrogate the specific structural mechanisms and design tradeoffs that enable particular functions (e.g. control of network dynamics). We are actively developing methods to fit and sample from these generative models at the scales needed to bring insight into the controllability properties of real-world connectomes.
Deep Networks for Computational Biology/Scientific Learning
 As scientific datasets grow in size and complexity, our models and methods must also grow to take advantage of the additional information. Modern AI tools, and deep networks in particular, have shown considerable performance benefits for hard prediction tasks from large, complex datasets. In neuroscience, we develop and apply AI models to decode behaviors, determine the computations occurring in the brain, and infer microscopic parameters from macroscopic observations. For example, using biophysically detailed simulations of the diversity of cortical neurons, we are training deep networks to infer the ionic conductances that produce experimentally accessible membrane potential observations. Preliminary results indicate that deep networks are particularly useful for addressing this inverse problem. We have additionally used deep networks to achieve state-of-the-art prediction of human speech production from ECoG recordings and as foundation models for brain machine interface decoding. In computational biology, we are developing and applying AI models to solve problems in genomics, molecular design, and molecular imaging. For example, modern AI has the potential to greatly accelerate protein design. Prior models have utilized purely protein sequence for this task, but protein function is most closely related to structure. We are training a foundation AI model jointly on protein sequence and structure data. Preliminary results indicate that the joint model is better able to relate sequence, structure, and function within its latent space, which improves the model’s ability to design useful proteins. In another AI thrust, we collaborate with the Berkeley Lab Molecular Foundry to improve structural imaging through deep learning for electron microscopy. Biological samples are notoriously fragile under electron beams, meaning that improving measurement efficiency can enhance imaging stability and the resulting knowledge of biological microstructure. We are developing a deep computer vision model for electron sensor readings, with the goal of improving single-electron localization and inter-electron segmentation.
As scientific datasets grow in size and complexity, our models and methods must also grow to take advantage of the additional information. Modern AI tools, and deep networks in particular, have shown considerable performance benefits for hard prediction tasks from large, complex datasets. In neuroscience, we develop and apply AI models to decode behaviors, determine the computations occurring in the brain, and infer microscopic parameters from macroscopic observations. For example, using biophysically detailed simulations of the diversity of cortical neurons, we are training deep networks to infer the ionic conductances that produce experimentally accessible membrane potential observations. Preliminary results indicate that deep networks are particularly useful for addressing this inverse problem. We have additionally used deep networks to achieve state-of-the-art prediction of human speech production from ECoG recordings and as foundation models for brain machine interface decoding. In computational biology, we are developing and applying AI models to solve problems in genomics, molecular design, and molecular imaging. For example, modern AI has the potential to greatly accelerate protein design. Prior models have utilized purely protein sequence for this task, but protein function is most closely related to structure. We are training a foundation AI model jointly on protein sequence and structure data. Preliminary results indicate that the joint model is better able to relate sequence, structure, and function within its latent space, which improves the model’s ability to design useful proteins. In another AI thrust, we collaborate with the Berkeley Lab Molecular Foundry to improve structural imaging through deep learning for electron microscopy. Biological samples are notoriously fragile under electron beams, meaning that improving measurement efficiency can enhance imaging stability and the resulting knowledge of biological microstructure. We are developing a deep computer vision model for electron sensor readings, with the goal of improving single-electron localization and inter-electron segmentation.
Statistical methods for system identification in neural population data
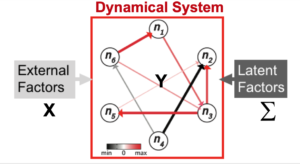 Neuroscience researchers often implicitly or explicitly interpret the output of their data analysis tools as reflecting the true state of nature. Therefore, neuroscience data analysis requires statistical machine learning algorithms that are simultaneously interpretable and predictive. By interpretable, we mean that inferred models yield insight into the physical processes that generated the data; by predictive, we mean that the model predicts the data with high accuracy. However, methods that achieve both are lacking. For example, while deep learning approaches can achieve remarkable predictive accuracy on extremely complicated data sets, extracting physically interpretable insights from the learned model remains a central challenge. Slightly more formally, algorithms for data driven discovery should be selective (only features that influence the response variable are selected), accurate (the estimated parameters in the model are as close to the “real” value as possible), predictive (allow prediction of the response variable), stable (return the same values on multiple runs) and scalable (able to return an answer in a reasonable amount of time on large data sets). We develop statistical machine learning methods that attempt to achieve all those goals in the context of systems identification in neural data.
Neuroscience researchers often implicitly or explicitly interpret the output of their data analysis tools as reflecting the true state of nature. Therefore, neuroscience data analysis requires statistical machine learning algorithms that are simultaneously interpretable and predictive. By interpretable, we mean that inferred models yield insight into the physical processes that generated the data; by predictive, we mean that the model predicts the data with high accuracy. However, methods that achieve both are lacking. For example, while deep learning approaches can achieve remarkable predictive accuracy on extremely complicated data sets, extracting physically interpretable insights from the learned model remains a central challenge. Slightly more formally, algorithms for data driven discovery should be selective (only features that influence the response variable are selected), accurate (the estimated parameters in the model are as close to the “real” value as possible), predictive (allow prediction of the response variable), stable (return the same values on multiple runs) and scalable (able to return an answer in a reasonable amount of time on large data sets). We develop statistical machine learning methods that attempt to achieve all those goals in the context of systems identification in neural data.
Quantifying spatial and dynamic complexity of neurons and behaviors
 Determining the complexity of the process that generated an observed time-series is a fundamental quantity that is directly related to the difficulty of learning a model of that process, as well as controlling it. Building off the seminal work of Bialek et al., 2001, we developed novel dimensionality reduction methods [Dynamical Components Analysis (DCA): linear-Gaussian model, Compressed Predictive Information Coding [CPIC]: non-linear, non-Gaussian model) that enable quantification of the complexity (defined as the predictive information between past and future observations) of the processes generating high-dimensional time-series data. We are applying these techniques to quantify the complexity of, e.g., vocal behaviors across species and conditions and single neuron input-output transformations towards developing the space, weight, and power (SWaP)- optimal neuronal unit for neuromorphic hardware.
Determining the complexity of the process that generated an observed time-series is a fundamental quantity that is directly related to the difficulty of learning a model of that process, as well as controlling it. Building off the seminal work of Bialek et al., 2001, we developed novel dimensionality reduction methods [Dynamical Components Analysis (DCA): linear-Gaussian model, Compressed Predictive Information Coding [CPIC]: non-linear, non-Gaussian model) that enable quantification of the complexity (defined as the predictive information between past and future observations) of the processes generating high-dimensional time-series data. We are applying these techniques to quantify the complexity of, e.g., vocal behaviors across species and conditions and single neuron input-output transformations towards developing the space, weight, and power (SWaP)- optimal neuronal unit for neuromorphic hardware.
Bridging spatiotemporal scales in the brain with multi-modal data and biophysical simulations

Biological systems are organized and function across a large range of spatial and temporal scales. One of the most complicated, multi-scale biological systems is the brain, in which myriad neurons are organized into microcircuits (‘columns’) that perform specific computations, but are simultaneously integrated into larger networks. Investigating the activity of individual neurons and small neuronal populations has yielded exquisite insight into microcircuit mechanisms of local computations, while macroscale measurements (e.g. fMRI) have revealed global processing of entire brain areas. Additionally, different frequency bands within the brains continuous electrical potential reflect different biological processes. For example, high-gamma (Hγ: 80-150Hz) activity is directly reflective of multi-unit neuronal spiking, while the lower frequency components (e.g. β: 15-30Hz) are thought to reflect synapto-dendritic currents. Thus, low frequencies might reflect inputs to neuronal populations, while the high-gamma band activity mainly reflects their spiking activity (outputs). How local neural processing is organized and coordinated in the context of broader neural networks, and how this is reflected in spiking activity and different frequency bands, is poorly understood. We combine uECoG with laminar polytrodes, optogenetic manipulations, and large-scale biophysically detailed simulations to bridge spatiotemporal scales in the nervous system. PLAY VIDEO of membrane potential in a large-scale biophysical simulation (note: this is slowed relatively to real-time. Movie curtesy of Burlen Loring).
Distributed representations of ethological auditory objects
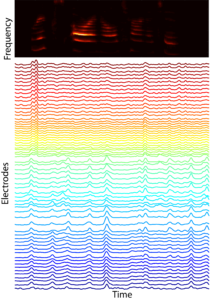 In many mammals, including humans, neural representations of complex sounds are distributed throughout large areas of cortex, including across primary auditory cortex (A1). The ability to discriminate between classes of sounds is critical to the survival of many species. For example, the ability of rats to distinguish between different avian sounds (e.g., hawk vs. crow) is a life-or-death capability. While the functional organization of A1 has been long investigated with simple stimuli, current receptive field models explain relatively small amounts (~15%) of response variation. This indicates that we do not understand what auditory features drive A1 neural responses. Concomitantly, most studies are focused on responses of relatively few neurons confined to a small number of cortical columns. While informative, such studies provide only limited insight into the distributed representation of ethologically relevant complex sounds. Indeed, we lack the broad sound stimulus set required to even begin addressing these issues. Thus, in order to advance our foundation for understanding representations of complex sounds we need a rich stimulus set along with spatially extended recording approaches to measure distributed representations of these sounds. We have collaborated in the design of novel mECoG devices that record neural activity spatially localized to ±200um (~the diameter of a cortical column), and which cover the entire A1. Our lab has pioneered concurrent laminar polytrode and mECoG recordings for simultaneous interrogation of local and distributed neural activity. We have constructed and continue to annotate a large database of natural sounds for playback during A1 recordings. Our recent analyses indicate that the structure of co-fluctuations amongst signals can provide insight into the nature of distributed representation. Together, these observations and tools motivate and enable us to test the hypothesis that A1 representations are optimized to classify complex, ethologically relevant sounds.PLAY VIDEO of neural responses to auditory stimuli, where each electrode is color coded by preferred frequency (note: this is at 1/4 speed).
In many mammals, including humans, neural representations of complex sounds are distributed throughout large areas of cortex, including across primary auditory cortex (A1). The ability to discriminate between classes of sounds is critical to the survival of many species. For example, the ability of rats to distinguish between different avian sounds (e.g., hawk vs. crow) is a life-or-death capability. While the functional organization of A1 has been long investigated with simple stimuli, current receptive field models explain relatively small amounts (~15%) of response variation. This indicates that we do not understand what auditory features drive A1 neural responses. Concomitantly, most studies are focused on responses of relatively few neurons confined to a small number of cortical columns. While informative, such studies provide only limited insight into the distributed representation of ethologically relevant complex sounds. Indeed, we lack the broad sound stimulus set required to even begin addressing these issues. Thus, in order to advance our foundation for understanding representations of complex sounds we need a rich stimulus set along with spatially extended recording approaches to measure distributed representations of these sounds. We have collaborated in the design of novel mECoG devices that record neural activity spatially localized to ±200um (~the diameter of a cortical column), and which cover the entire A1. Our lab has pioneered concurrent laminar polytrode and mECoG recordings for simultaneous interrogation of local and distributed neural activity. We have constructed and continue to annotate a large database of natural sounds for playback during A1 recordings. Our recent analyses indicate that the structure of co-fluctuations amongst signals can provide insight into the nature of distributed representation. Together, these observations and tools motivate and enable us to test the hypothesis that A1 representations are optimized to classify complex, ethologically relevant sounds.PLAY VIDEO of neural responses to auditory stimuli, where each electrode is color coded by preferred frequency (note: this is at 1/4 speed).
Coordination of distributed cortical networks producing complex movements

The generation of simple skilled behaviors, such as reaching to grasp objects, requires the rapid and precise coordination of many body parts (e.g. shoulder, elbow, wrist, digits). As the representation of different body parts is spatially distributed across sensorimotor cortex (SMC), the coordination required of skilled behavior implies similar coordination of SMC generating movements. Despite many experiments in SMC, we have limited understanding of the functional organization and coordination of neuronal populations that produce skilled behaviors. Recent studies have emphasized the importance of local neural population dynamics; however, examination of larger scale network dynamics is rare. Furthermore, linking dynamics to principles of representation has proven challenging. Compared to our understanding of motor networks, our knowledge of sensory processing is more developed. The concept of sparsity has proven very useful to understand computations in sensory systems, and has recently been applied to spatial patterns of hippocampus field potential recordings; however, the role of sparseness in motor regions is not understood. We conjecture that primary motor regions transform sparse independent representations of afferent messages from premotor areas into dense coordinated efferent signals for driving motor actuators. Extending our recent findings in humans/rodents, we will understand brain functions across multiple spatiotemporal scales, examine how distributed representations are dynamically coordinated, and test hypotheses on the role of sparsity in cortical computations. We have developed a novel, 3 d.o.f robot to enable 3D reach tasks in rats with video tracking. PLAY VIDEO of rat reaching to handle.
Recent Comments